|
Situation one favors only one tail of the distribution. Question: Why might observed and expected phenotype frequencies differ? Imagine the following scenarios where natural selection is at work. Significantly different, the population is out of HWE. If observed and expected genotype frequencies are Square p to get p 2 and multiply 2*p*q to get the observed heterozygous Aa genotype frequency. Then, take the square root of q 2 to get q, and then subtract q from 1 to get p. Count up the aa types and you have the observed q 2. If the alleles are dominant and recessive, we can't visually tell the homozygous AA from the heterozygous Aa genotypes (both are tall), so it's best to start with the homozygous recessive (short) aa individuals. Tip: If the alleles are codominant, each phenotype is distinct (you can distinguish between tall, medium and short) and your job is easier. In other words, the observed frequency of A = 100%(AA) + 50%(Aa) 
Value, you can of course just subtract it from 1 (100%) to get the Half of the Aa individuals (the a half) plus all of aa individuals. The observed frequency of allele a is therefore However, in a population of genotypes AA, Aa and aa, the observedįrequency of allele A equals the sum of all of the AA genotype plus 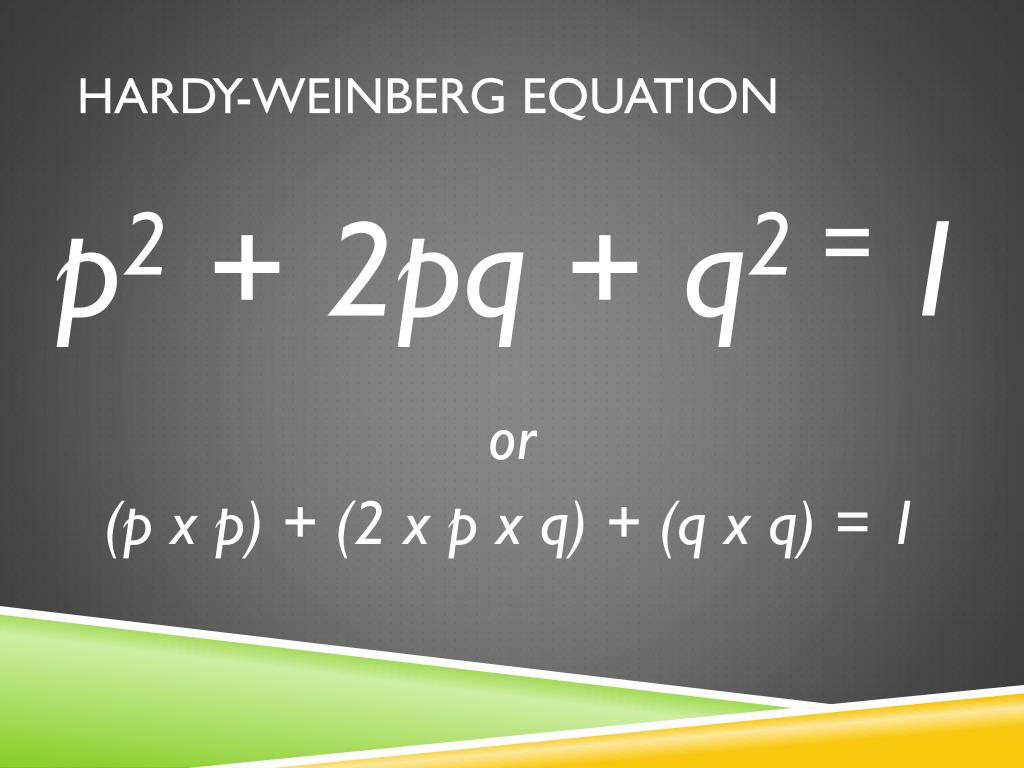
In a real world population, we can only see phenotypes, not genotypes or alleles. Next, let's look at the real world situation so we can compare.Ĭalculating Observed Allele and Genotype Frequencies: Red represents the frequency of the AA or A 1A 1 genotype, green is the Aa or A 1A 2 genotype, and blue is the aa or A 2A 2 genotype.Īll of the above has to do with the allele and genotype frequencies we would expect to see. For values of p from 0 to 1, in intervals of 0.1, here's what we get: Let's take a look at some graphs of this to make it a little easier to see. The expected frequencies of the genotypes are therefore: In other words, p 2 + pq + pq + q 2 = 1, orġ00%. Multiply the allele frequencies to the get the probability of each genotype. Allele A or A 1 has a frequency of p, and allele a or A 2 has a frequency of q. The expected genotype frequencies of the two alleles are calculated as shown. If we know the allele frequencies, we can predict the genotype frequencies. (Because there are only two possibilities and they have to add up to 100%, p + q = 1.) They will have frequencies p and q in a population. 
Don't worry for now whether the alleles are dominant and recessive or co-dominant. Let's say that A or A 1= tall, and a or A 2= short. These alleles might be A and a, or A 1 and A 2. In the simplest possible situation we have a single gene with only two alleles. one or more of the above) is going on, andĬalculating Expected Allele and Genotype frequencies: Significantĭifferences between the observed and expected frequencies indicate They allow us to calculate an expected allele frequency. Genetic Drift is not occurring (drift is less likely in populations of large size)Īlthough these assumptions are rarely true in the natural world,.Generation if the following assumptions are met: Will remain in Hardy-Weinberg Equilibrium (HWE) from generation to Hardy-Weinberg Equilibrium Hardy-WeinbergĪllele frequencies (or percentages, if you prefer) in a population
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
